Summary
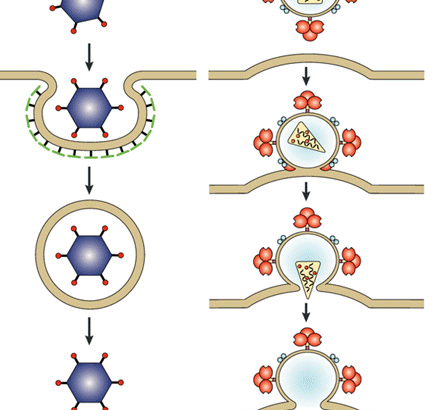
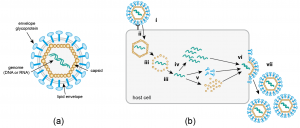
Viruses are intracellular parasites that hijack cellular machinery for their own replication. Therefore, a mandatory step in the virus life cycle is the delivery of the viral genome into the cell. Entry of enveloped viruses into cells (i.e., viruses with a lipid envelope) use a two-step procedure to release their genetic material into the cell: (i) they first bind to specific surface receptors on the target cell membrane and then, ( ii) fuse the viral and cellular membranes. This latter step may occur at the cell surface or after internalization of the viral particle by endocytosis or some other pathway (eg, macropinocytosis).
Remarkably, the virus-cell membrane fusion process follows essentially the same intermediate steps as other membrane fusions that occur, for example, in vesicle fusion at nerve synapses or cell-cell fusion in yeast mating. . Specialized viral proteins, fusogenic, promote fusion of the virus-cell membrane. Viral fusogenic undergo drastic structural rearrangements during fusion, releasing the energy needed to overcome the repulsive forces that prevent spontaneous fusion of the two membranes.
Keywords: class I fusion protein, class II fusion protein, class III fusion protein, enveloped virus, fusion pore, glycoprotein, membrane, membrane fusion intermediate, post-entry events, viral fusogen, virus entry
The Mechanical Properties Of The Influenza Virus Change According To Host Signals
Influenza virus passes through acidic endosomes, an essential step for shedding and infection. While residing in low pH conditions, protons pass through the viral M2 ion channel into the lumen of the virus and there separate the viral ribonucleoprotein (vRNP) cores from the M1 protein by inducing a conformational change in M1. This is consistent with the observation that the M1 protein has multiple functions in the virion and occurs in a ribbon or coiled structure.
Proton entry into the virion lumen then softens the viral envelope, most likely by dissociating vRNPs from the inner side of the envelope. Evidence for this notion was obtained by atomic force microscopy measurements with native virions and “bald” particles lacking viral glycoproteins. The latter result implies that low pH-induced changes in viral glycoproteins do not contribute significantly to envelope softening under low pH conditions.
Host Mechanics Control TheDeveloping Genome Of Adenovirus And Influenza Virus
Many viruses, including adenoviruses, infect postmitotic cells and replicate in the nucleus. Adenovirus deposits its linear DNA genome, but not the viral capsid, in the nucleus, as click-chemically tagged incoming virus genomes at single-molecule resolution recently demonstrated. The virus uses the molecular motor kinesin-1 to separate the genome from the capsid. This occurs on the cytoplasmic face of the nuclear pore complex (NPC), where the major capsid protein of the virus couples to the nucleoporin Nup214. The kinesin-associated 1,2 light chain binds to another virus capsid protein (protein IX), and the motor domain on the heavy chain is activated by binding to Nup358 of the filaments of the cytoplasmic pore complex.
Microtubules are located proximal to nuclear pore complexes because they have a binding site in the distal domain of Nup358. This quinary complex of viruses, two Nups and two motor components, then executes virus disruption and coincidentally part of the nuclear pore complex as well. The viral DNA (presumably in a complex with the viral protein VII) is then imported into the nucleus with the help of cellular transport factors, such as importins and transportin. However, the nuclear import process of viral DNA is not perfectly accurate and a variable fraction of the incoming viral DNA is misdelivered to the cytoplasm. This raises questions about innate immune recognition of cytosolic viral DNA and possible viral antagonism.
In addition to adenovirus, shedding of the influenza virus genome depends on cellular motor proteins, in this case, dynein and myosin. Under conditions of low pH in the endosome, the influenza virus alters the conformation of the glycoprotein hemagglutinin, leading to exposure of the hydrophobic fusion peptide and fusion of the viral membrane with the limiting endosomal membrane. Cytosolic RNPs dissociate from the membrane fusion site by a process that mimics the formation and removal of offenders.
This involves the unanchored ubiquitin-binding domain (ZnF-UBP) and dynein-binding domains of histone deacetylase 6 (HDAC6), as well as microtubule- and actin-based motors. Since the ionic milieu in endosomal compartments is subject to regulation (late but not early endosomes contain high concentrations of potassium ions and low concentrations of sodium ions, for example), additional signals acting on the virus can be anticipated. endosomal. In fact, the high concentrations of potassium ions together with the low pH in late endosomes prime the influenza A virus to separate vRNPs well before fusion.
Conclusions And Perspectives
This assay has attempted to integrate the physical properties of individual virus particles with mechanical or chemical signals from host cells. Physical and structural virology will continue to elucidate novel and exciting features of individual virus particles and show that they are important for infection, virus transmission, or vaccination against viral diseases. However, the full power and importance of these features are only manifested if the physical and structural features are integrated into the context of cellular or immunological mechanisms.
This approach considers the environment in which viruses have been selected in the course of evolution. Furthermore, it is important to note that viruses act as a swarm of particles and genetic elements, and a single virus particle rarely manages to infect a single cell. Rather, more often than not, virus particles cooperate or compete during infection. This notion gives rise to the emerging field of pathogen coinfection: for example, virus-virus and virus-bacteria coinfections.